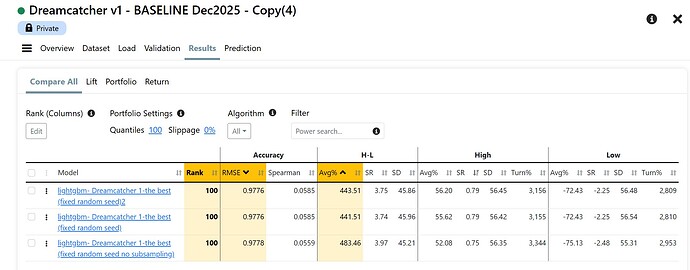
Just thinking out loud and troubleshooting with you. I have not actually tried much of this.
I think it helps to separate the different sources of randomness in tree-based models. In practice there are three:
-
Row sampling (bootstrapping / subsampling)
-
Feature sampling (column subsampling)
-
Split randomness (specific to ExtraTrees)
For Random Forests, randomness comes mainly from (1) and (2): deterministic splits, but random rows and features.
For ExtraTrees, the key additional source is (3): randomized split thresholds, even when using the full dataset (no randomization of rows or features).
That split randomness is usually sufficient for ExtraTrees to control variance, which means bootstrapping and column subsampling are usually unnecessary for ExtraTreesRegressor. Turning those off typically has little impact on performance.
So in effect, with ExtraTreesRegressor you can remove two sources of randomness (row and feature sampling) while relying on a third (randomized splits). The advantage is that the remaining randomness operates on a fixed dataset, rather than being amplified by small changes in row ordering or indexing as would be the case with a random forest that uses bootstrapping and/or features subsampling.
Maybe you have already tried this and whether this fully resolves the behavior you’re seeing, I can’t say for sure — but in principle it should reduce run-to-run variability while preserving the core behavior of ExtraTrees.
I’m not suggesting ExtraTrees as a replacement for LightGBM here — but it might be useful as a diagnostic test. Effective because the whole data set is always used with ExtraTreesRegressor (with bootstrapping/subsamplng and feature subsampling turned off). If removing row and feature sampling collapses the variation across copied AI factors, that would strongly suggest those mechanisms are amplifying small upstream differences. If it doesn’t, then the randomness is likely entering earlier in the pipeline.
Again just troubleshooting, I asked ChatGPT and it focused on this too (part of its answer):
The most important insight (this is the punchline)
seed only guarantees reproducibility for repeated runs on the same realized dataset.
Creating a new AI Factor implicitly creates a new dataset realization.
Or my words for that: Its a different array and maybe using the entire data set rather than slicing it and dicing in using subsampling with minimize the effect of the changes.